Environmental microbiology is the study of the relationships between microorganisms and the environment, including the composition, structure and physiology of microbial communities. The environment not only consists of the soil, water, air, sediment, and rocks covering the planet, but also includes the animals and plants that inhabit these areas and microbes that exist in artificial environments such as bioreactors. Research in environmental microbiology is highly diverse and employs multiple cross-disciplinary approaches. A number of UNH faculty are engaged in various collaborative research programs addressing fundamental environmental issues addressing how organisms respond to environmental changes.
W. Kelley Thomas — Marine biodiversity and environmental change
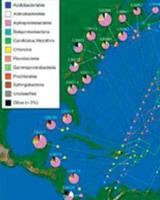
The Thomas lab has developed high throughput approaches to investigate biodiversity with a particular focus on the meiofauna (very small eukaryotes). Our current focus is on marine sediments which house vast amounts of the global biodiversity and are potentially impacted by environmental change and anthropogenic events like the Deepwater Horizon Oil Spill.
W. Kelley Thomas
Gregg Hall, Room 448
(603) 862-2470
kelley.thomas@unh.edu
Louis Tisa — Environmental signals regulating microbial relationships
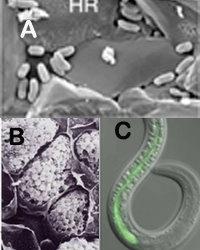
The Tisa lab uses genome-wide and genome-guided approaches towards addressing environmental issues. Our work emphasizes interactions between microbes and: plants, nematodes, rock, and other microbes. One area of research centers on Frankia, a beneficial bacterial symbiont of actinorhizal plants. We are investigating plant-microbe communication signals that result in this symbiotic relationship. In collaboration with Feixia Chu at UNH, we employ proteomic approaches to identify key proteins in this communication process that may assist in developing potential bioremediation strategies for heavy-metal or pollutant-contaminated land.
A second major research focus centers on insect biological control agents. The Photorhabdus-Heterorhabditis symbiosis is a well-known, commercially available biological control agent, and our lab is investigating broadening its use to include other arthropods. The bacterium maintains two distinct lifestyles: as a nematode symbiont and as an insect pathogen. The bacteria are voracious pathogens to a myriad of insect larvae and generate essential growth factors for the nematode. Nematode reproduction and development are an obligate requirement for the bacteria. Our lab is studying these bacteria to better understand the roles of motility, surface properties, and biofilm formation in insect pathogenesis and nematode symbiosis, and how these processes are regulated during the life cycle of Photorhabdus.
Louis Tisa
Rudman Hall, Room 289
(603) 862-2442
louis.tisa@unh.edu
Cheryl Whistler — Host-microbe interactions

The Whistler lab has several areas of active inquiry in environmental microbiology. One prominent area of research deals with environmental factors affecting host association in both pathogenic and symbiotic bacteria in the genus Vibrio. We apply a broad range of molecular genetic and genomic techniques to define similarities and differences in symbionts and pathogens. Our experimental models include an elegant mutualistic symbiosis between Vibrio fischeri and its squid host, as well as an environmentally transmitted pathogen (Vibrio parahaemolyticus) for which virulence is elusive and poorly defined. Another research area explores the difference between the planktonic (free-living) and host associated lifestyles for both these Vibrios by comparing strains from a broad biogeographical distribution and with different association capacities; we seek to better define how they have evolved and adapted to different lifestyles. Finally, in collaboration with Drs. Steve Jones and Vaughn Cooper at UNH, we are examining the population structure of human pathogenic Vibrio species within the context of their native microbial communities in different ecological niches (including native oysters in the Great Bay Estuary; see photo). We are applying sophisticated methodologies including MLSA analysis of Vibrio populations and analysis of the oyster metagenome to generate specific hypotheses about how biotic (ecology) and abiotic (e.g., climate change) factors influence the emergence of specific bacterial community members, especially the emergence of pathogenic bacteria.
Cheryl Whistler
Rudman Hall, Room 208
(603) 862-2359
cheryl.whistler@unh.edu
Serita Frey — Soil microbial ecology

Serita Frey’s research specifically examines how anthropogenic stressors (e.g., climate change, nitrogen deposition, agricultural management, invasive species) affect the composition and diversity of soil microbial communities and microbial-mediated carbon and nitrogen cycles. We work at the interface between ecosystem science, microbial ecology and soil science, combining microbiological methods with stable isotope analysis and a variety of soil physical and chemical fractionation techniques to examine structure-function linkages. We work in a variety of ecosystems, including temperate forests, freshwater wetlands, and agroecosystems.
Serita Frey
James Hall, Room 172
(603) 862-3880
serita.frey@unh.edu
Stephen Jones — Ecology of pathogenic bacteria in aquatic ecosystems

My focus is on Vibrio species that cause disease in humans and associate with shellfish, and fecal-borne bacteria that pollute surface waters. Research focuses on their sources, their fate and impacts, and strategies to eliminate them as problems. New approaches include genomic studies of microbial communities and practical strategies to ensure shellfish safety.
Steve Jones
Jackson Estuarine Laboratory
(603) 862-5124
stephen.jones@unh.edu
Jessica Ernakovich — Microbial Communities and Elemental Transformation

The Ernakovich lab seeks to understand the factors that structure natural microbial communities and the impact of that organization on their function, particularly their role in elemental transformations. We do this with a mixture of tools, from DNA and RNA sequencing to use of stable isotopes. We have worked in a variety of environments, from the Arctic tundra to the Outback in Australia. Current focal areas are New Hampshire agricultural systems and thawing permafrost in Alaska and Sweden.
Jessica Ernakovich
James Hall 176 (office)
Rudman Hall 132 (lab)
(603) 862-2216
jessica.ernakovich@unh.edu
Twitter: @JessErnakovich
Jeff Garnas — Ecology and evolution of insects and microbes in forest systems

The Garnas lab focuses on understanding how insects, microbes, and climate interact to affect the health of forests. Projects of greatest relevance to the MCBS program include 1) understanding how variation in tree mycobiomes influence the severity of decline in American beech infected with beech bark disease; 2) characterizing population structure and host-switching dynamics, gene flow, and selection in Neonectria pathogens on beech and non-beech hosts; and 3) using high throughput sequencing and digital droplet PCR to examine emerging diseases of white pine as a function of climate, tree stress, and overlapping pathogen distributions. Our field, lab, and molecular work is rooted in population, community, and evolutionary ecological principles and often focuses on the consequences of biological invasion and climate change for forest ecosystems.
Jeff Garnas
James Hall, Room 162
(603) 862-2094
jeff.garnas@unh.edu